Want to share your content on R-bloggers? Click here if you have a blog, or here if you don't.
Overview
Building on the R6-based unifiedml package introduced last week, this post covers mlS3 — a new R package that provides a unified, consistent S3 interface for training and generating predictions from a range of popular machine learning models. Instead of adapting to the unique API conventions of each underlying package (you'll still want to consult their documentation for parameter details), mlS3 wraps them behind a shared wrap_*() / predict() pattern.
What You'll Learn
- How to install and load
mlS3(currently available from GitHub) - How to apply a consistent API across multiple ML algorithms for both classification and regression tasks
Models Covered
| Wrapper | Underlying Package | Task(s) |
|---|---|---|
wrap_glmnet() |
glmnet — generalized linear models |
Classification, Regression |
wrap_lightgbm() |
lightgbm — gradient boosting |
Classification, Regression |
wrap_ranger() |
ranger — random forest |
Classification, Regression |
wrap_svm() |
e1071 — support vector machines |
Classification, Regression |
wrap_caret() |
caret — 200+ models |
Classification, Regression |
Datasets Used
iris— binary and multiclass classification (setosa vs. versicolor; all three species)mtcars— regression to predict miles per gallon (mpg)
Key Design Principle
Every model in mlS3 follows the same two-step workflow:
mod <- wrap_*(X_train, y_train, ...) # Train pred <- predict(mod, newx = X_test, ...) # Predict
This uniform interface makes it straightforward to swap algorithms and compare results without restructuring your pipeline.
Prerequisites
- R with the following packages:
remotes,caret,randomForest,ggplot2 mlS3installed from GitHub viaremotes::install_github("Techtonique/mlS3")
Code
Installing Packages
install.packages(c("remotes"))
install.packages(c("caret"))
install.packages(c("randomForest"))
remotes::install_github("Techtonique/mlS3")
Predefined Wrappers
# Classification
library(mlS3)
# =============================================================================
# Classification examples (no leakage)
# =============================================================================
set.seed(123)
# --- Binary classification: iris setosa vs versicolor ---
iris_bin <- iris[iris$Species != "virginica", ]
X_bin <- iris_bin[, 1:4]
y_bin <- droplevels(iris_bin$Species)
# Split into train/test
idx_bin <- sample(nrow(X_bin), 0.7 * nrow(X_bin))
X_bin_train <- X_bin[idx_bin, ]
y_bin_train <- y_bin[idx_bin]
X_bin_test <- X_bin[-idx_bin, ]
y_bin_test <- y_bin[-idx_bin]
# glmnet
mod <- wrap_glmnet(X_bin_train, y_bin_train, family = "binomial")
pred_bin_glmnet <- predict(mod, newx = X_bin_test, type = "class")
acc_glmnet <- mean(pred_bin_glmnet == y_bin_test)
cat("Accuracy (glmnet): ", acc_glmnet, "\n")
# --- Multiclass classification: iris all species ---
X_multi <- iris[, 1:4]
y_multi <- iris$Species
# Split into train/test
idx_multi <- sample(nrow(X_multi), 0.7 * nrow(X_multi))
X_multi_train <- X_multi[idx_multi, ]
y_multi_train <- y_multi[idx_multi]
X_multi_test <- X_multi[-idx_multi, ]
y_multi_test <- y_multi[-idx_multi]
# lightgbm
mod <- wrap_lightgbm(X_multi_train, y_multi_train,
params = list(objective = "multiclass",
num_class = 3, verbose = -1),
nrounds = 150)
pred_multi_lightgbm <- predict(mod, newx = X_multi_test, type = "class")
acc_lightgbm <- mean(pred_multi_lightgbm == y_multi_test)
# ranger
mod <- wrap_ranger(X_multi_train, y_multi_train, num.trees = 100L)
pred_multi_ranger <- predict(mod, newx = X_multi_test, type = "class")
acc_ranger <- mean(pred_multi_ranger == y_multi_test)
# svm
mod <- wrap_svm(X_multi_train, y_multi_train, kernel = "radial")
pred_multi_svm <- predict(mod, newx = X_multi_test, type = "class")
acc_svm <- mean(pred_multi_svm == y_multi_test)
cat("Accuracy (lightgbm): ", acc_lightgbm, "\n")
cat("Accuracy (ranger): ", acc_ranger, "\n")
cat("Accuracy (svm): ", acc_svm, "\n")
# =============================================================================
# Regression examples (mtcars)
# =============================================================================
X_reg <- mtcars[, -1]
y_reg <- mtcars$mpg
# Split into train/test
set.seed(123)
idx_reg <- sample(nrow(X_reg), 0.7 * nrow(X_reg))
X_reg_train <- X_reg[idx_reg, ]; y_reg_train <- y_reg[idx_reg]
X_reg_test <- X_reg[-idx_reg, ]; y_reg_test <- y_reg[-idx_reg]
# lightgbm
mod <- wrap_lightgbm(X_reg_train, y_reg_train,
params = list(objective = "regression", verbose = -1),
nrounds = 50)
pred_reg_lightgbm <- predict(mod, newx = X_reg_test)
rmse_lightgbm <- sqrt(mean((pred_reg_lightgbm - y_reg_test)^2))
# glmnet
mod <- wrap_glmnet(X_reg_train, y_reg_train, alpha = 0)
pred_reg_glmnet <- predict(mod, newx = X_reg_test)
rmse_glmnet <- sqrt(mean((pred_reg_glmnet - y_reg_test)^2))
# svm
mod <- wrap_svm(X_reg_train, y_reg_train)
pred_reg_svm <- predict(mod, newx = X_reg_test)
rmse_svm <- sqrt(mean((pred_reg_svm - y_reg_test)^2))
# ranger
mod <- wrap_ranger(X_reg_train, y_reg_train, num.trees = 100L)
pred_reg_ranger <- predict(mod, newx = X_reg_test)
rmse_ranger <- sqrt(mean((pred_reg_ranger - y_reg_test)^2))
cat("RMSE (lightgbm): ", rmse_lightgbm, "\n")
cat("RMSE (glmnet): ", rmse_glmnet, "\n")
cat("RMSE (svm): ", rmse_svm, "\n")
cat("RMSE (ranger): ", rmse_ranger, "\n")
Accuracy (glmnet): 1
Accuracy (lightgbm): 0.2444444 # I'm probably doing something wrong here
Accuracy (ranger): 0.9333333
Accuracy (svm): 0.9333333
RMSE (lightgbm): 4.713678
RMSE (glmnet): 2.972557
RMSE (svm): 2.275837
RMSE (ranger): 2.067692
The caret Wrapper
This section requires the caret and randomForest packages. The full list of models available through wrap_caret() can be found here: https://topepo.github.io/caret/available-models.html
library(mlS3)
library(caret)
# =============================================================================
# Regression with mtcars dataset
# =============================================================================
data(mtcars)
X_reg <- mtcars[, -1] # All columns except mpg
y_reg <- mtcars$mpg # Target variable
# Split into train/test
set.seed(123)
idx_reg <- sample(nrow(X_reg), 0.7 * nrow(X_reg))
X_reg_train <- X_reg[idx_reg, ]
y_reg_train <- y_reg[idx_reg]
X_reg_test <- X_reg[-idx_reg, ]
y_reg_test <- y_reg[-idx_reg]
# ----------------------------------------------------------------------------
# Example 1: Random Forest Regression
# ----------------------------------------------------------------------------
cat("\n=== Example 1: Random Forest Regression ===\n")
mod_rf <- wrap_caret(X_reg_train, y_reg_train,
method = "rf",
mtry = 3) # Variables sampled at each split
print(mod_rf)
# Predictions and evaluation
pred_rf <- predict(mod_rf, newx = X_reg_test)
rmse_rf <- sqrt(mean((pred_rf - y_reg_test)^2))
r2_rf <- 1 - sum((y_reg_test - pred_rf)^2) / sum((y_reg_test - mean(y_reg_test))^2)
cat("RMSE:", round(rmse_rf, 3), "\n")
cat("R-squared:", round(r2_rf, 3), "\n")
=== Example 1: Random Forest Regression ===
$model
Random Forest
22 samples
10 predictors
No pre-processing
Resampling: None
$task
[1] "regression"
$method
[1] "rf"
$parameters
$parameters$mtry
[1] 3
attr(,"class")
[1] "wrap_caret"
RMSE: 2.007
R-squared: 0.681
library(ggplot2)
df <- data.frame(
pred = pred_rf,
actual = y_reg_test
)
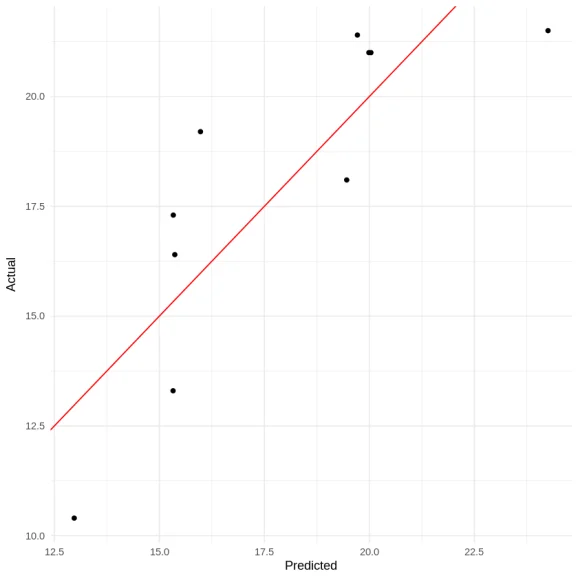
ggplot(df, aes(x = pred, y = actual)) +
geom_point() +
geom_abline(slope = 1, intercept = 0, color = "red") +
theme_minimal() +
labs(x = "Predicted", y = "Actual")
R-bloggers.com offers daily e-mail updates about R news and tutorials on learning R and many other topics. Click here if you're looking to post or find an R/data-science job.
Want to share your content on R-bloggers? Click here if you have a blog, or here if you don't.
mlS3 — A Unified S3 Machine Learning Interface in R


