Want to share your content on R-bloggers? Click here if you have a blog, or here if you don't.
This article is cross-posted on rOpenSci and R-Consortium blogs.
For more than two decades, the Bioconductor project has served as a cornerstone of the R ecosystem — delivering peer-reviewed, high-quality tools for bioinformatics and computational biology. Its curated repository model, rigorous review standards, and tightly coordinated release cycles have made it one of the most trusted distribution channels in scientific computing.
Yet sustaining a project of this age and scale comes at a cost. Legacy build systems, bespoke tooling, and organically grown workflows accumulate technical debt that becomes increasingly expensive to maintain. To address this, Bioconductor is partnering with R-universe to incrementally modernize its infrastructure — preserving the project's governance, quality standards, and established processes while reducing operational overhead. The collaboration is mutually beneficial: as Bioconductor adopts R-universe components, it also helps shape and refine the platform's feature set to meet the demands of a large, complex scientific community.
The partnership aligns with R-universe's broader mandate as an R Consortium Infrastructure Steering Committee (ISC) top-level project: to provide modern, open, and reusable infrastructure that strengthens the R ecosystem and supports reviewed package repositories like rOpenSci and Bioconductor.
Two Universes: Release and Development
Bioconductor maintains two distinct package branches:
- A release branch for stable, production-ready packages
- A devel branch for active development and the upcoming release cycle
To mirror this structure, two dedicated R-universe instances are now in operation:
- Development branch: https://bioc.r-universe.dev
- Release branch: https://bioc-release.r-universe.dev
Both universes integrate directly with Bioconductor's existing Git infrastructure and provide continuous builds across both branches. Through the R-universe dashboard, package maintainers and users can:
- Inspect cross-platform check results
- Review extended BiocCheck diagnostics
- Monitor build logs and dependency graphs
- Explore rich package metadata and activity metrics
- Access binary packages for Windows, macOS, and Linux
The result is a familiar yet modern interface for Bioconductor contributors — consistent with what users increasingly expect from contemporary R package infrastructure.
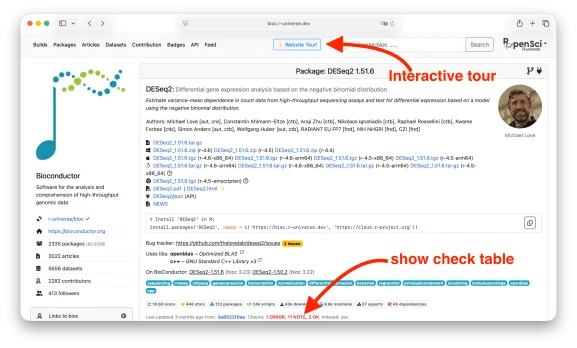
Individual package pages are available at https://bioc.r-universe.dev/{pkgname}. For example, https://bioc.r-universe.dev/DESeq2 provides a full overview of the DESeq2 package:
First-time visitors are encouraged to click the "Website Tour" button for a guided overview of the most important features — it takes only one to two minutes.
Technical Documentation for Bioconductor Maintainers
Comprehensive technical documentation for R-universe is available at https://docs.r-universe.dev. A dedicated section has been created specifically for Bioconductor developers, covering the most relevant topics for getting started: https://docs.r-universe.dev/bioconductor/
The documentation will expand as the collaboration matures and new components are introduced. The goal is to give Bioconductor maintainers a clear, practical reference for understanding how R-universe fits into their development workflow — without disrupting the practices that have made Bioconductor a trusted institution within the R community.
Looking Ahead
Adopting new infrastructure always requires adjustment. Bioconductor developers will likely need to adapt some workflows, and invest time in becoming familiar with updated package checks, build diagnostics, and binary distribution mechanisms.
The longer-term payoff, however, is substantial. By converging on shared, actively maintained infrastructure, the Bioconductor project stands to benefit from continuous improvements developed for the broader R ecosystem. A modern CI-based platform will give developers better tooling and free the core team to concentrate on community coordination and quality control — rather than sustaining aging infrastructure. The shared visibility provided by R-universe also stands to make Bioconductor software more discoverable and accessible to the wider R community.
Both teams look forward to deepening this partnership and working alongside the Bioconductor community to ensure the next generation of infrastructure serves the project well for years to come.
© 2025 Bioconductor. Content published under Creative Commons CC-BY-4.0 License for text and BSD 3-Clause License for code. | R-Bloggers
R-bloggers.com offers daily e-mail updates about R news and tutorials on learning R and many other topics. Click here if you're looking to post or find an R/data-science job.
Want to share your content on R-bloggers? Click here if you have a blog, or here if you don't.